This version improves the user interface in various places, implements a new search, and adds nodes for querying OME data and XML/Text/JSON processing.
User interface improvements
Parameter editor
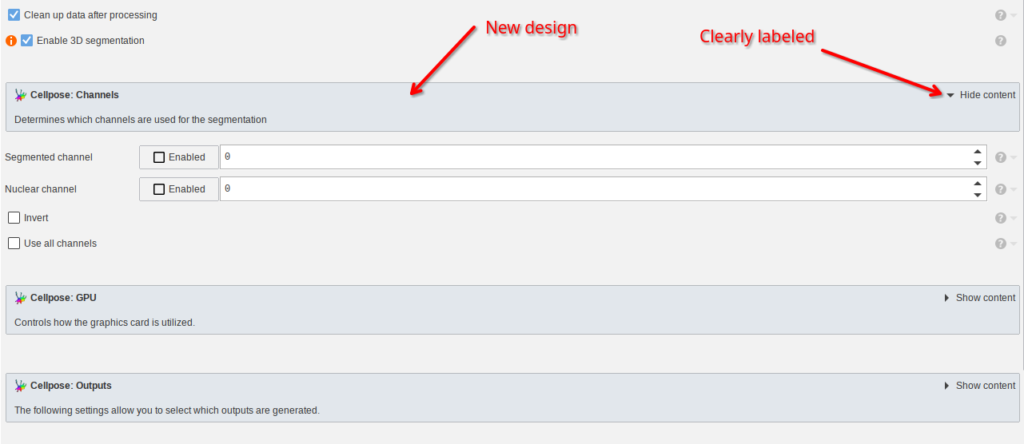
The design of all parameter editors was refined:
- The header bar is now visually separated from the parameters
- The collapse/expand button is now clearly described
Node search & algorithm finder
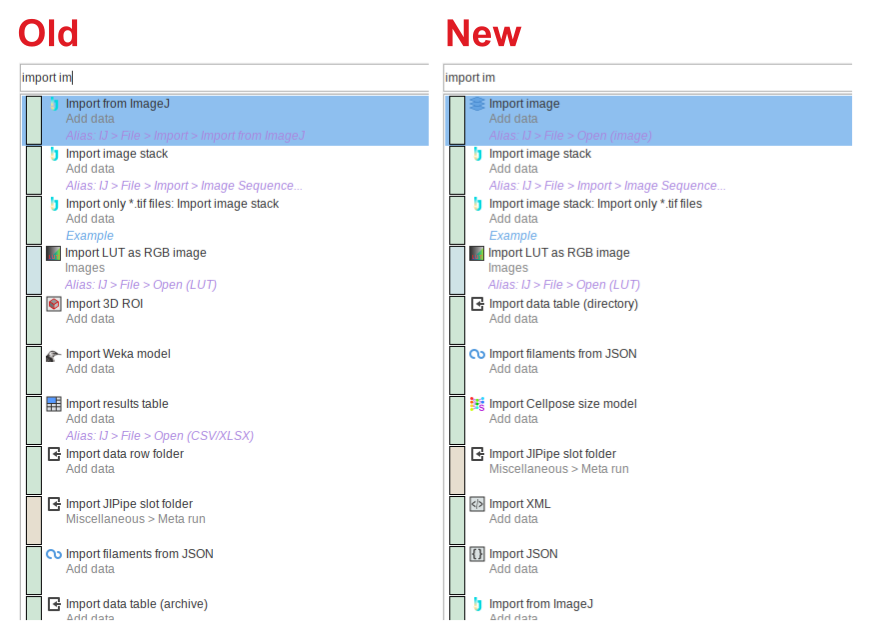
The node search implemented in previous versions produced inconsistent results, as the search bar, algorithm finder, and node list utilized different algorithms for applying filtering and ranking nodes.
A new unified node search was implemented that generates consistent results for all search operations and additionally implements a new algorithm that generates better results.
The algorithm finder was redesigned to be more simple and faster. The new interface uses the new node search.
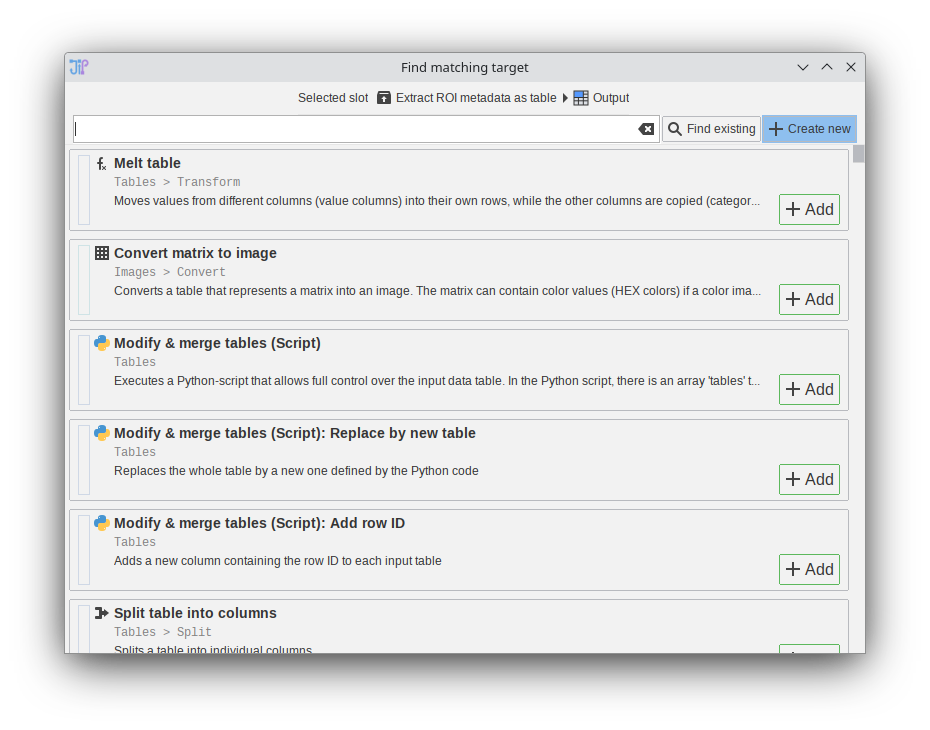
Tip: if you want to quickly add the first entry in the list, press enter on the keyboard to trigger the “Add” button.
Tip: you can now double-click a slot to trigger the algorithm finder
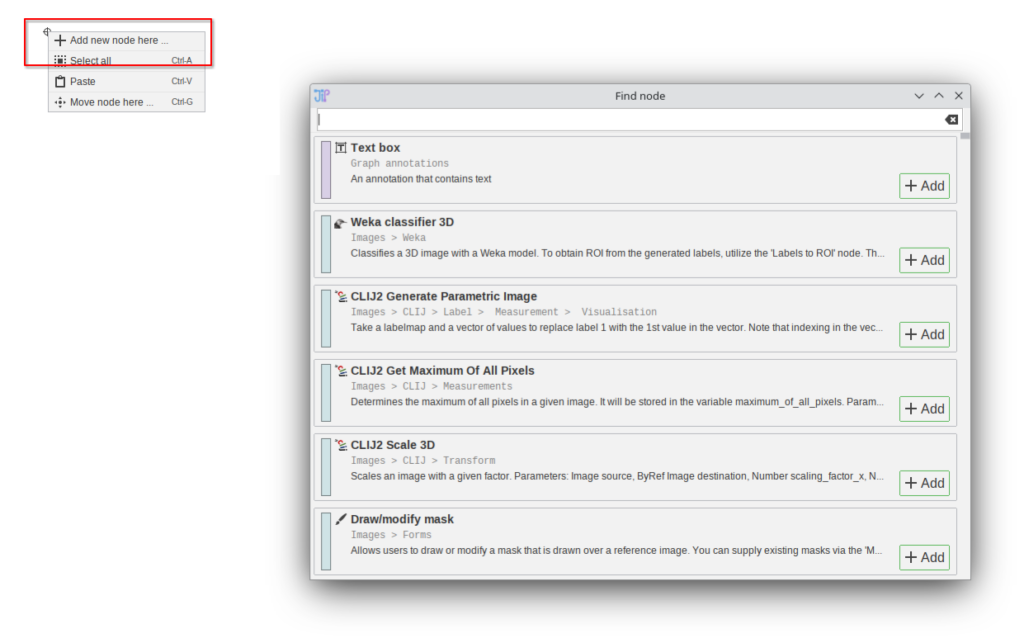
A standalone algorithm finder (without the need for an existing slot) can be triggered by right-clicking the graph and selecting “Add new node here …”
Errors/issues
The new JIPipe version comes with a redesigned error panel that allows users to jump to node that caused the error.
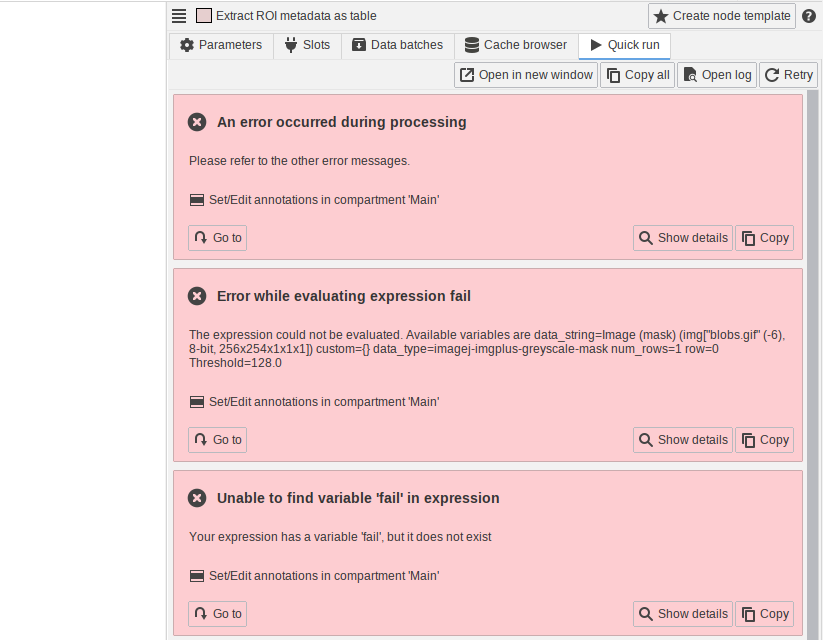
Data tables
Data tables displaying the current cache can now be right-clicked to open a context menu
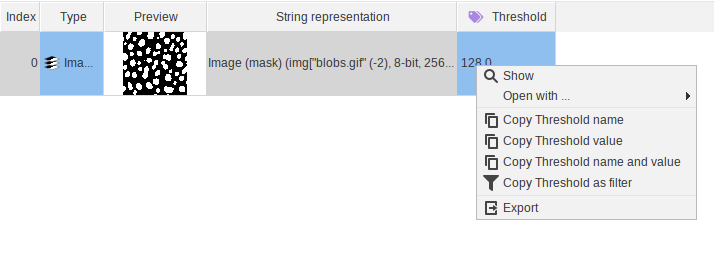
Expression editor
The toolbar of the expression editor was redesigned to be easier understandable. An additional option was introduced that allows the selection of a path.
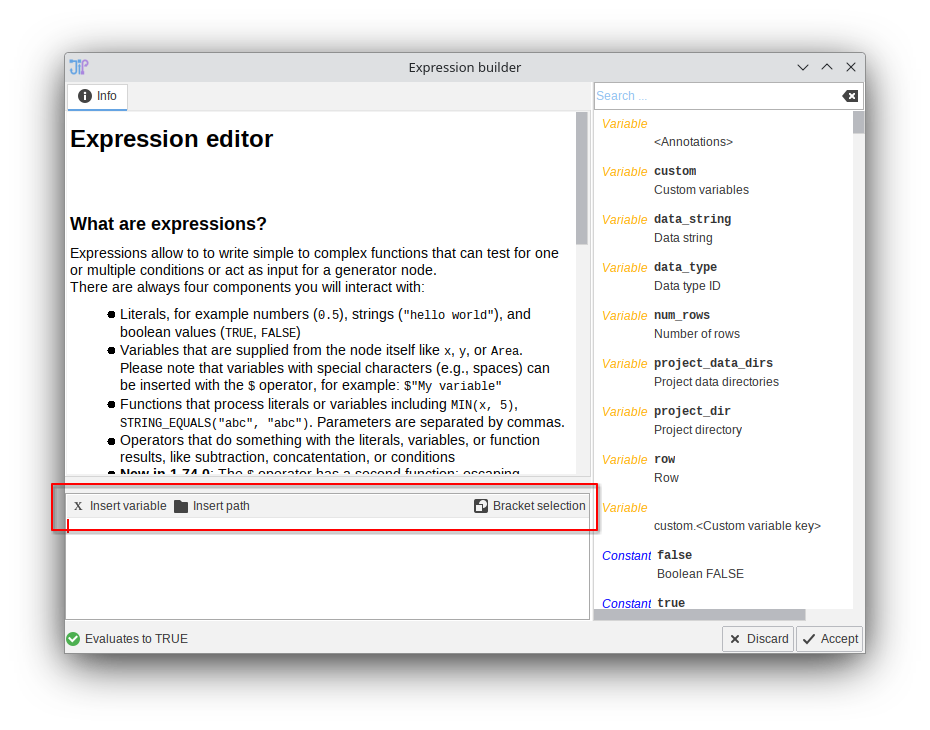
Script editor
The script editor component was improved and now only allows one instance at the same time.
Additionally, it is now possible to edit scripts in an external editor (changes to the external file are automatically copied into the JIPipe parameter)
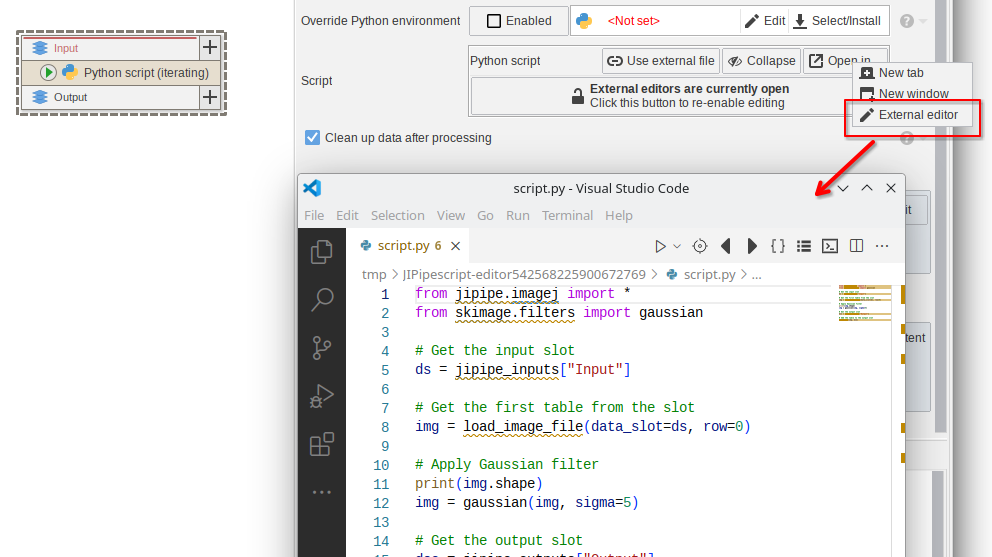
OME image processing
It is now easier to extract information from OME images
- New node “Extract OME ROI” (was only accessible via convert before)
- New node “Extract OME XML” (was only accessible via convert before)
- New node “OME metadata as table”
- New node “Annotate with OME metadata”
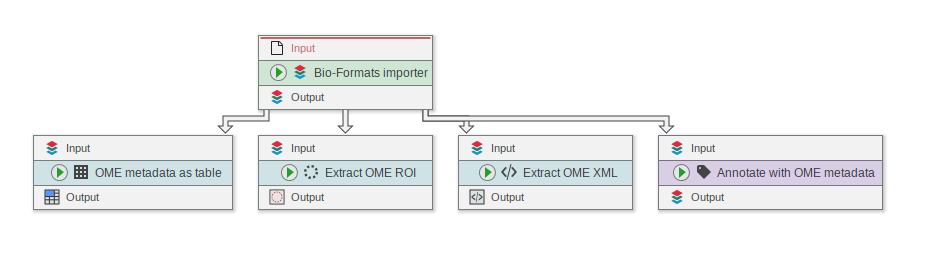
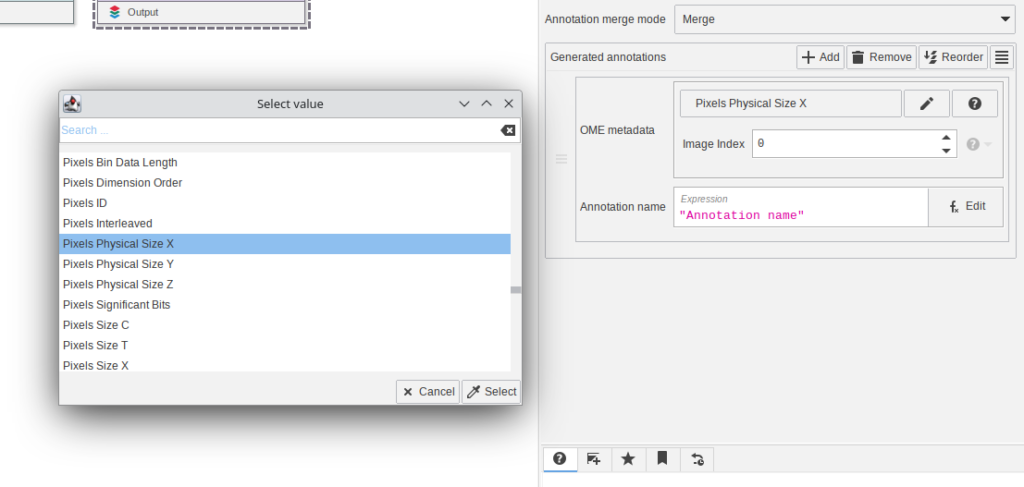
“OME metadata as table” and “Annotate with OME metadata” provide easily accessible user interfaces to select OME metadata items.
Data export
Most data export nodes were redesigned* to utilize a single “File path” attribute instead of handling the output directory and file name independently. Instead of providing many confusing options for automatically generating file names, a default name is provided via the auto_file_name variable. Customization is offered by the expression language.
* Older exporter nodes were marked as deprecated, but continue to work.
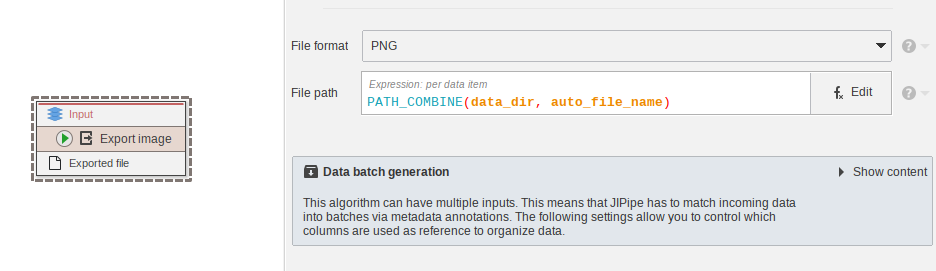
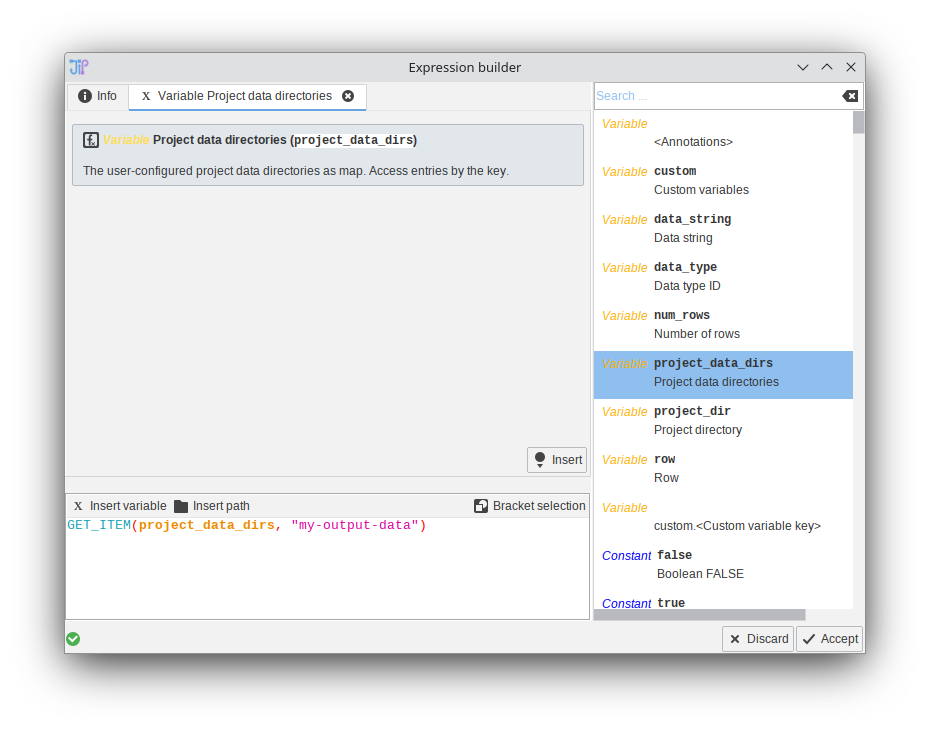
The expression allows the storage of files in various locations, including the default data directory that is relative to the project run output (data_dir), the project directory (project_dir), fully custom locations (just type in the path as string), and the newly introduced project-wide user directories.
Project-wide user data directories
The new JIPipe version introduces a new project-wide setting that allows to configure a set of directories that can be accessed inside various nodes.
The setting can be accessed as following:
- Project > Project overview > User directories
- or: Project > Project settings > User directories
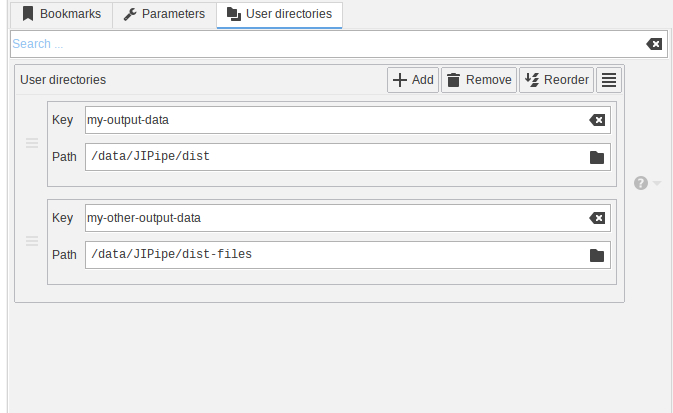
The interface allows to create new entries that have a “Key” and “Path” setting. The key is used to lookup directories in expressions that offer the project_data_dirs variable.
Python
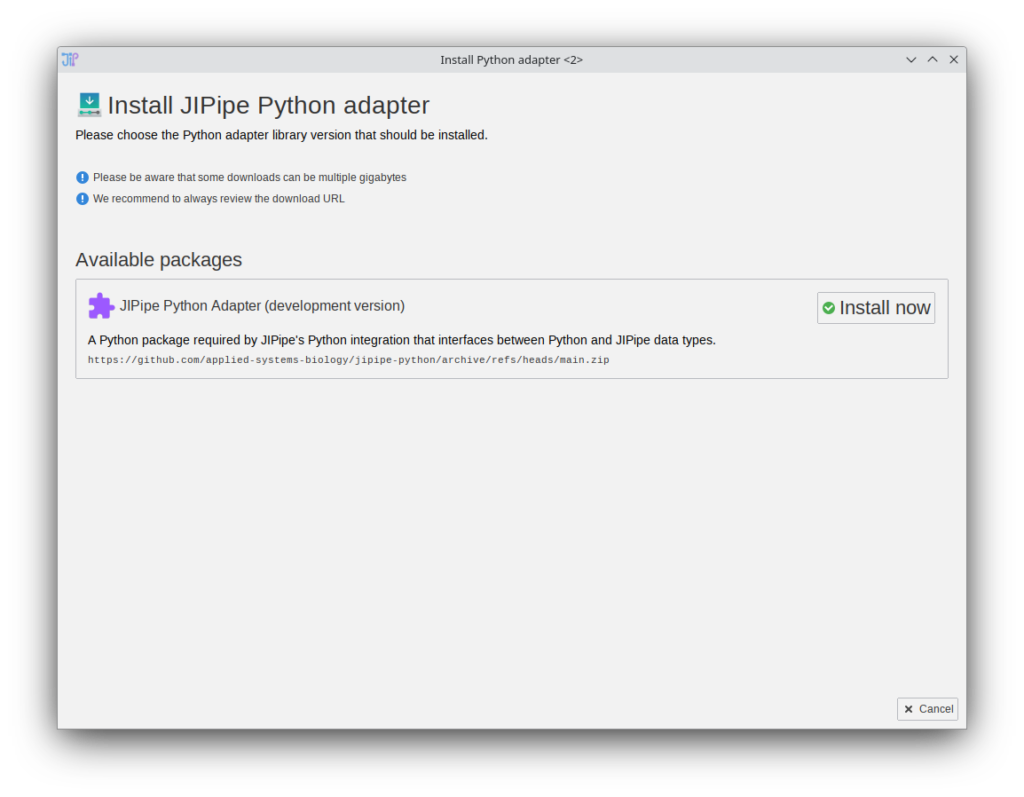
JIPipe uses a Python adapter library (JIPipe-Python) to communicate with Python scripts. In previous JIPipe versions this adapter was provided with the JIPipe plugin, which makes it hard to apply updates. The setup process was converted into an EasyInstaller environment. Additionally, JIPipe now checks for updates for the Python adapter during the startup (this can be changed by navigating to Project > Application settings > Extensions > Python integration (adapter) and unchecking “Automatically check for updates”
Existing JIPipe users will be prompted to install the Python adapter once.
XML/Text/JSON processing
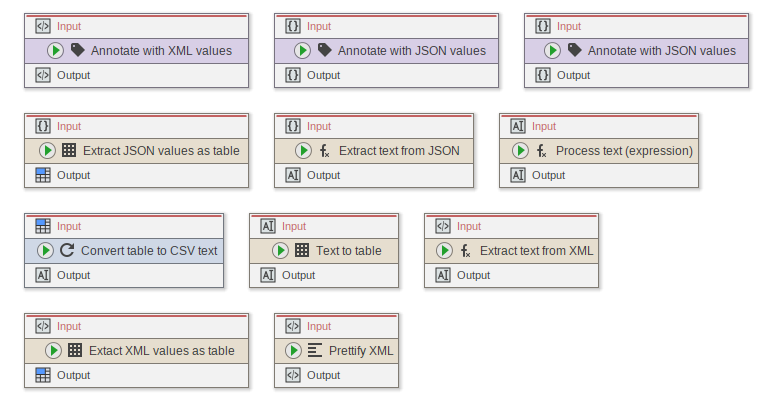
JIPipe now comes with nodes that allow to modify/extract information from XML, text, and JSON data
- New node “Annotate with text values” (using an expression)
- New node “Annotate with JSON values” (using JSON path)
- New node “Annotate with XML values” (using XPath)
- New node “Extract JSON values as table”
- New node “Extract text from JSON”
- New node “Process text (Expression)”
- New node “Convert table to CSV text”
- New node “Text to table”
- New node “Extract text from XML”
- New node “Extract XML values as table”
- New node “Prettify XML”
Project report
JIPipe can now generate a project report that contains metadata, dependencies, citations (including ones inferred from utilized nodes/extensions/plugins), an overview of all utilized nodes, and a generated text description of the pipeline.
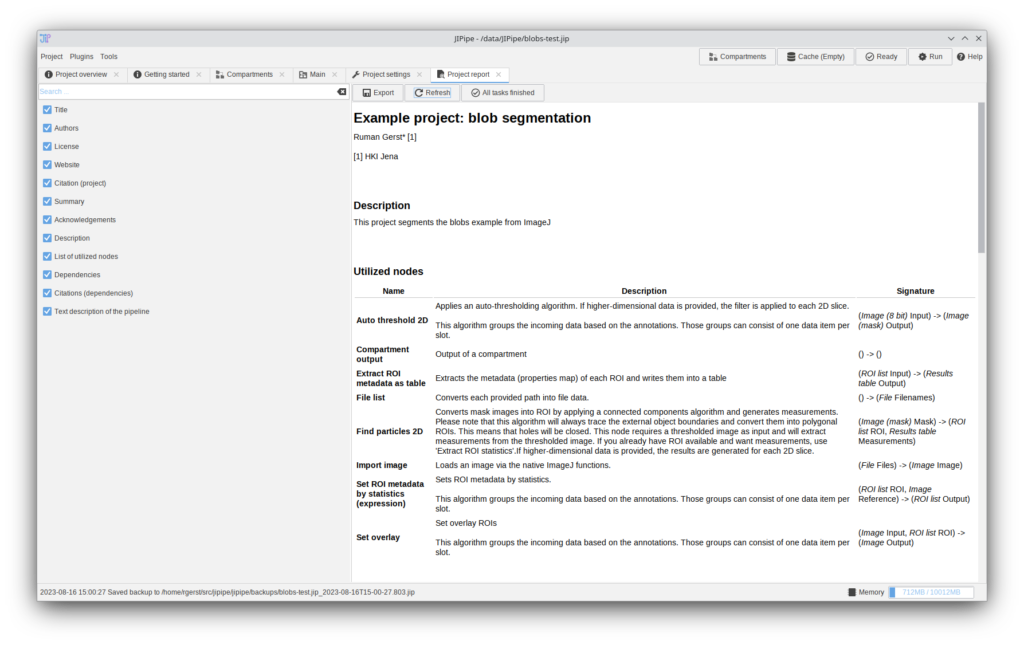
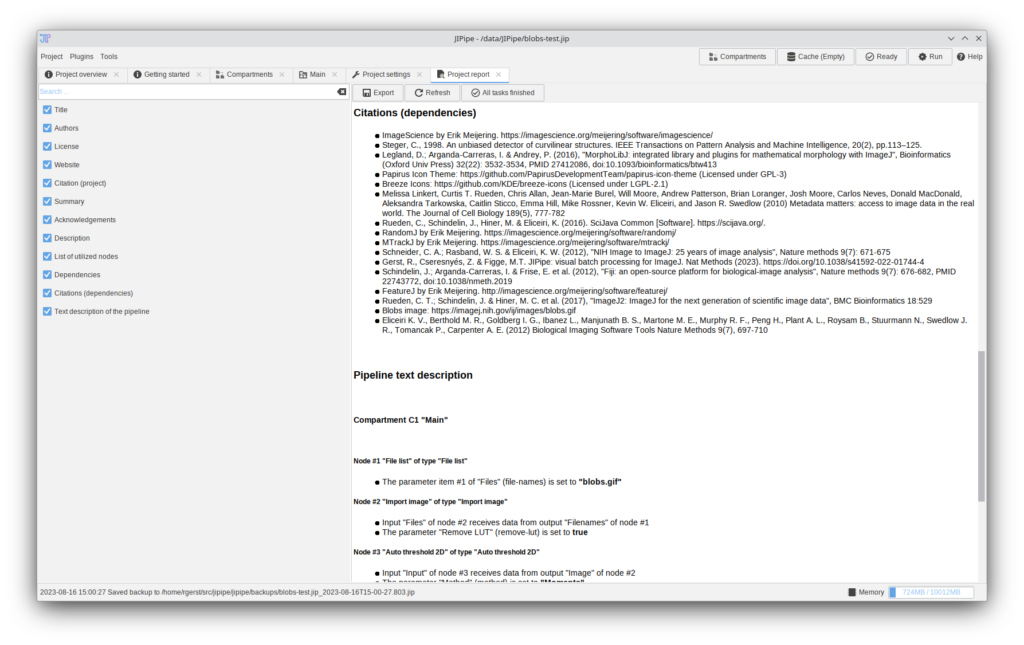
The report shown in the screenshot can be found here:
Expressions
- Added constant: new_line (\n)
- New function: IS_FINITE(number)
- New function: IS_INFINITE(number)
- New function: IS_NAN(number)
- GET_PARENT_DIRECTORY now has a second parameter to select the Nth parent directory
- Support for running Python (Jython) and Javascript code within expressions using the JYTHON() and JS() functions
API
- Extensions now can provide node templates via resources
- Extensions can now provide examples via direct instantiation in postprocess()
- The error API was fully redesigned
Small improvements
- Find particles 2D: added missing option to output composite ROI (holes etc)
- Improved “Data batches” UX
- Improved naming of various nodes
- UX: better icons for various elements
- Annotate with data: control over how annotations are merged
- Generate image from math expression: now has annotation input
- “Modify path”, “Set/edit annotations”, and “Set single annotation” now have access to project data directories
- CLIJ now avoids keeping GPU memory allocated by default (more stable in our experience)
- Examples were added for various nodes
Bugfixes
- Pivot table node was not registered
- Table transform operations are now properly categorized
- Fix bug in rewire tool
- Nodes could move to the mouse cursor on loading a pipeline
- Locked nodes could be deleted and copied
- Tables could not display XML/HTML values (now properly escaped)
- Drag & drop into the graph areas was not functional (cursor stuck at one location)
- UX: dragging a selection over a locked node did not yield a selection marquee
- The skipping of incomplete data batches was not logged
- Performance: Cellpose ROI extraction is now only executed if required
- R: now works again with annotations
- BioFormats importer: was not always able to extract series ID
- Split into training/test (90:10) was generating image outputs
